I am happy to introduce the padr package, which is now available on CRAN. If you frequently work with data containing a timestamp, especially automatically created data, you might find this package helpful. It solves two problems that you can be confronted with when preparing datetime data for analysis. First, data is often recorded on too low a level for your analysis. For instance the timestamp records the moment up to the second, where you want to do the analysis on an hourly level. Second, when no events toke place there are typically no data records. This is sensible from a storage perspective, but often unhelpful for analyzing the data. When calculating a moving average for example, you want missing observations to have the value 0. You don’t want them to be lacking from your set.
Aggregate data with thicken
When your data is on too low a level, you need to aggregate it to a higher level. thicken creates this higher level. Lets say we want to graph the expenses at a coffee place and we have a data set containing a timestamp and the amount spent.
library(padr)
library(dplyr)
coffee %>% thicken(interval = "day")
## time_stamp amount time_stamp_day
## 1 2016-07-07 09:11:21 3.14 2016-07-07
## 2 2016-07-07 09:46:48 2.98 2016-07-07
## 3 2016-07-09 13:25:17 4.11 2016-07-09
## 4 2016-07-10 10:45:11 3.14 2016-07-10
So we fed thicken the data frame and it did a few things for us. First, it figured out what the datetime variable in the dataset was. Then it noticed the interval of the dataset, which is “second” in this case. Next, it mapped each timestamp to the corresponding “day”, which is the desired interval of the output. Finally, it added the mapping as a new column to our data frame. We can now easily aggregate to this new variable, for instance using the dplyr functions group_by and summarise.
coffee_day <- coffee %>%
thicken(interval = "day") %>%
group_by(time_stamp_day) %>%
summarise(total = sum(amount))
coffee_day
## # A tibble: 3 <U+00D7> 2
## time_stamp_day total
## <date> <dbl>
## 1 2016-07-07 6.12
## 2 2016-07-09 4.11
## 3 2016-07-10 3.14
Fill gaps with pad
When we would make a line graph of coffee_day, it would skip over 2016-07-08, when there was no visit. This is not what we wanted. Since the expenses at this day were 0, we would like a record here indicating there were no puchases. We now invoke the pad function to insert this record.
coffee_full <- coffee_day %>% pad
coffee_full
## # A tibble: 4 <U+00D7> 2
## time_stamp_day total
## <date> <dbl>
## 1 2016-07-07 6.12
## 2 2016-07-08 NA
## 3 2016-07-09 4.11
## 4 2016-07-10 3.14
Like thicken, the function figured out what the datetime variable was, and that it has the interval “day”. It then noticed that an observation is missing from this interval, and it created a record for it. The other variable, total, got a missing value inserted for this record.
Use fill_ for missing values
So pad will always leave us with a data frame with missing values for the inserted records. How we want to fill this missings depends on the data at hand. Sometimes you would like to carry the last value forward, you can then use tidyr::fill. In this case we would like to fill the NA with 0, for which we use fill_by_value. Other functions available in padr are fill_by_function and fill_by_prevalent.
library(ggplot2)
coffee_full %>% fill_by_value(total) %>%
ggplot(aes(time_stamp_day, total)) + geom_line()
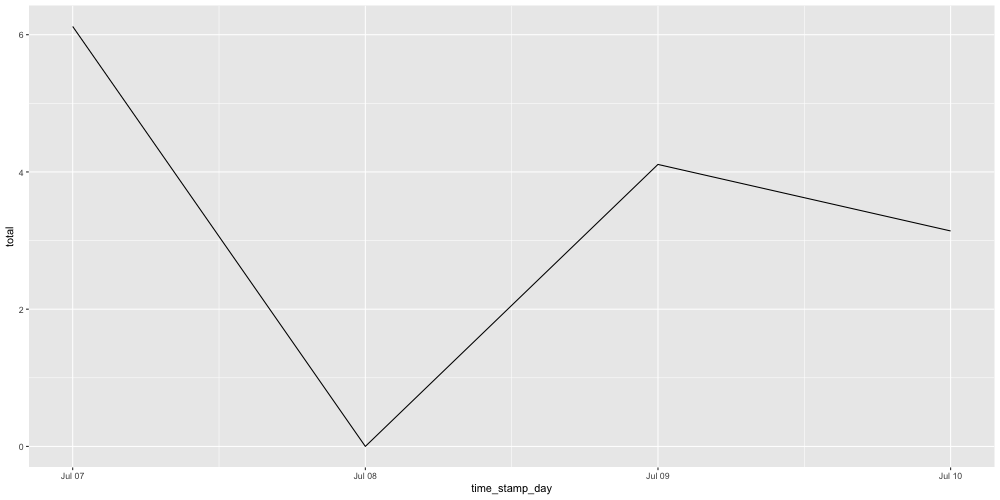
So here we have our plot, and here we have our quick overview of padr. A more thorough introduction can be found in vignette("padr"). Here I also get into more detail on the meaning of “interval” within the package. Feedback is very welcome, either send me an email, open an issue on github, or do a pull request. The code is maintained here. Finally, I’d like to thank Barret Schloerke for his contributions at making this package CRAN ready.
